Illumina presented several key updates at a technology roadmap session on Thursday. There's been extensive coverage of this in press, Twitter and by analysts, which I will not try to link to systematically to all though I can't omit links to a trio of friends: Julianna LeMeuix's GEN piece, James Hadfield's nice writeup and a Twitter user Varro Analytics collated multiple analysis reports. Also I'll note that my colleague BG was taking notes in Slack as fast or faster than I was and I've borrowed some of the images she snagged.
After much waffling back-and-forth, I've decided to split this into two reports: one on the new short read sequencing chemistry and hardware with a separate one on the Illumina's is-it-or-is-it-not synthetic read technology. A bit later, some pontificating on how the market now stacks up in my eyes.
Big Boxes Sequencers Are In...
The core rollout is the NovaSeq Xplus, a successor to NovaSeq for the highest throughput operations. The $1.25M instrument sports fully independent flowcells and claims to deliver $2/Gb data for a $200 genome. Instruments won't be shopped until late Q1 with Illumina expecting demand to outstrip supply through all of Q2 and into Q3.
The box ain't small -- it looks an awful lot like a Sequel IIe. Nava Whiteford has noted it weighs in at nearly three quarters of a metric ton, not quite the likes of an RS II or a Helicos.
A single flowcell model will ship sometime next year as well, the NovaSeq X, selling for about $400K [sadly, I originally wrote $40K - what's a factor of 10-fold $ between friends?] lower. I always wonder on these how many of the chopped down models Illumina actually sells versus how many it gets someone to commit to and then successfully upsells them to the full box.
Chemistry X has been officially branded "XLEAP-SBS". Illumina touts a raft of improvements -- more stable chemistry, heavily engineered polymerase, faster cycle times, lower phasing and so forth. Manufacturing of the reagents and flowcells taking place in San Diego, the UK and Singapore, with each site supporting a different region and giving resilience to Illumina's ability to supply reagents - and the validation plus testing of the platform has already involved 4000 flowcells.
A geeky detail I didn't spot: which way does it read the indexes? Forward Strand or Reverse Complement workflow? It's required knowledge for actually setting up the sample sheet correctly and Illumina has flipped between the two schemes roughly every other platform rollout.
The new box has a lot of fancy new optics -- 4X the pixels, 10X faster scanning, 2X the data transfer rate, higher resolving power (sub 50nm) - to go with the fancy new chemistry. This is apparently a key reason for the new chemistry not appearing on the legacy NovaSeq. The NextSeq 1000/2000 instruments do have sufficient optics and a P4 flowcell for these using the new chemistry will also show up -- but not until 2024.
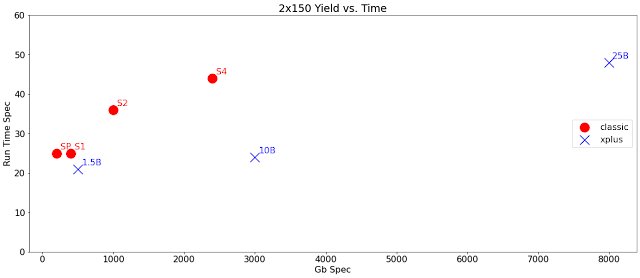
I've plotted out below the yield per runtime for 2x150 modes on the three announced NovaSeq Xplus flowcells and the four existing NovaSeq flowcells. Illumina isn't giving code names to the new ones -- each one is named after its nominal number of clusters. In 2x150 mode, the 1.5B delivers more than an SP or S1 in few hours less and the 10B delivers more than an S2 or S4 with substantial time savings -- as in 20 hours faster than S4.
An annoyance amidst all the announcements is that it is often difficult to clearly match up claims to specific products. Illumina I fear likes a certain level of fog -- you aren't supposed to immediately grok that the 24 hour turnaround claim and 16Tb run claims don't apply to the same flowcell. As a another example, is the $2/gigabase going to be the pricing for all three flowcells in 2x150 mode or just the 25B?
...Big Box Supplies Are Out
A ritual I feel a bit guilty never participating in is the surprisingly involved affair that goes on every time an Illumina shipment comes in. Some supplies are on dry ice, some on cold packs and everything is in large boxes, so all the packages must be broken down and packed into the appropriate cold storage. With the new chemistry, Illumina will be shipping NovaSeq X series supplies at ambient temperature, eliminating the need for dry ice and its large packaging as well as cold packs. The cartridges themselves are half the size, saving space in storage refrigerators. The cartridges are even made of a "biodegradable" plastic. though too often such plastics require a lot of encouragement to biodegrade.
This was a huge hit with the crowd -- this isn't the most flashy tech but it really matters practically plus it means less waste and carbon footprint. Getting dry ice out of the supply chain is very timely, given reports of CO2 shortages. Plus dry ice is classified as a hazardous substance -- for Warp Drive I was trained in proper shipping and disposal (not down the drain if you like your pipes!!). Eliminating this is also a huge win for less developed countries with limited cold chain facilities -- plus, as one London Calling attendee once described to me, corrupt port officials really know they have you over a barrel if your dry ice is sublimating away -- the bribe demands are more insistent and for larger amounts.
Though the optics on NovaSeq apparently can't support the higher density flowcells in the NovaSeq X series, I'm sure a lot of labs running NovaSeq would love to see the unpacking parties go away, which would be enabled by the new reagents.
DRAGEN: An M&A Big Win!
Illumina has often enhanced their platform through mergers and acquisition. The entire sequencing-by-synthesis platform was an acquisition of Solexa and the Nextera technology came in from Epicentre. M&A doesn't always pan out -- Illumina's acquisition of Advanced Liquid Logic begat the doomed NeoPrep library prep robot and seemingly nothing else and the Moleculo acquisition produced another product fizzle.
Edico will go down as another huge win for Illumina, yielding the DRAGEN technology. It isn't a panacea, but for Illumina's core business of enabling human genome SNP calling DRAGEN has demonstrated in many head-to-head competitions that it is top notch. NovaSeq X series instruments will have DRAGEN built in, following the pattern of the NovaSeq 1000/2000 set. Indeed, I think it would be very surprising if any future Illumina hardware doesn't have DRAGEN onboard
Despite the huge size of the NovaSeq X case, the original prototype apparently left no room for the compute! Further miniaturization -- and presumably some of the wetware being compressed - allowed a very powerful multicore package to be onboard. This handles basecalling, demuxing of barcodes and human alignment and variation calling. Presumably other genomes can be swapped in, though with the DRAGEN FPGA architecture you don't want to frequently swap -- there's a time penalty. In Illumina's hands DRAGEN's variant calling is nearly as good as DeepVariant and somewhat better than GATK -- so if you are comparing platforms for variant calling performance and using Illumina+GATK that isn't an appropriate benchmark anymore.
Illumina should be expected to continue to hammer on the DRAGEN quality and speed -- for any other platform it is a fair question to ask what downstream analyses have been subcontracted to the sequencer rather than requiring off-instrument compute and likely a bioinformatics team to supervise the operation.
Illumina Listens the Same Market Research as Everyone Else
A number of features of the new instrument resemble features of new players such as Ultima, Element and Singular. Unless Illumina employs some very effective clandestine operatives, it's reasonable to assume that everybody is hearing the same pleas from the existing user base and any market researchers. There's eight independent lanes on a flowcell (presumably only the 25B; the pattern has been for more lanes to yield more clusters), so less barcoding is absolutely required. The two flowcells can be run completely independently, in contrast to NovaSeq where a second flowcell cannot be started while the first flowcell is clustering.
An interesting corollary is that NovaSeq Xplus is not being presented as an exome / enrichment killer. Illumina gave some gaudy number for the total exomes that could be run on one 25B flowcell. It will be interesting to see the degree to which the market decides that with $200 short read genomes the extra hassle of capture isn't really worth it, particularly since those non-captured regions will likely yield interesting biology in the future.
No Long Reads the Size of Short Reads
The announced kits include 2x50, 2x100 and 2x150 for the 1.5B and 10B flowcells but only 2x150 for the 25B flowcell. So that says that Illumina isn't seeing an opportunity for very high numbers of shorter reads to support counting applications whereas Element and Singular do -- but of course the smaller sequencers are supporting around 1B reads per flowcell so really could be compared to the 1.5B flowcell
Illumina had been suggesting previously that Chemistry X (now XLEAP-SBS) would support longer reads, but there is no sign of it in the initial offerings -- no equivalent of the 2x250 mode on the SP flowcell of the NovaSeq classic. The longer reads are a bit niche, but valuable niches such as protein engineering. There are some other potential limits -- the lab folks say that patterned flowcells don't like inserts much above 500 basepairs.
No Love For the Low Rungs of the Product Ladder
The only instrument announcement not about NovaSeq X or NextSeq 1000/2000 was the announcement of a registered diagnostic version of the NovaSeq 6000. Clearly NovaSeq platform will be around for a bunch longer, but at some point will Illumina start phasing out non-Dx versions of the reagents -- making any holdouts pay a premium price
So iSeq and MiniSeq continue their pattern of getting no updates and the venerable MiSeq platform was shut out as well. Certainly these lack the necessary optics to support the high density XLEAP-SBS flowcells, but as noted above the benefits of ambient shipping have no resolution requirements.
Even if XLEAP-SBS only enables longer reads at existing density, that would seem to be marketable --2x250 and 2x300 are very popular on MiSeq. Note that on these older platforms using bridge amplification for clustering, very long inserts can be tolerated. There were limited papers describing clustering at over 1 kilobase, though these yielded (if I recall correctly) oversized clusters that degraded the density which could be supported. Still, for specialized applications such as MHC haplotyping the idea of 2x400 reads on 800 basepair amplicons could get some people very excited.
Sign-Off
As always, what did I miss (other than what I've deferred to a separate post on Infinity / Illumina Complete Long Reads)? What did I botch? Pour on the comments and brickbats please!

4 comments:
What do you think about the accuracy of new Chemistry? Although Illumina claimed that the new chemistry has a higher accuracy, but the q-score spec did not seem to have improved.
"A single flowcell model will ship sometime next year as well, the NovaSeq X, selling for about $40K lower."
Hi Keith, this should be $400K lower, not 40K. Or rather it should be 1.3M - 0.985M = $315K?
I believe it was stated that initially the X Plus would not have independent running flow cells. When launched they will be required to run together. A later development will allow for the independence.
Hi Keith, wonderful summery in your very entertaining style. Thanks for that! But, without being picky... 1,250,000 - 985,000 = 265,000. Isn`t it?
Post a Comment