Before 2015 ends, I'd like to tie up two loose threads. In doing so, I'll deviate slightly from my usual pattern and publish two posts in a day; I could have lumped them together but instead I'll split. First up, a belated explanation, prompted by a comment, of my mention of issues with the MiSeq 2x300 reagents and a bit more on my confusion with regard to bootstrap values.
Back in my item on BGI shelving the Revolocity platform, I remarked in passing that Illumina has had recent problems with the 2x300 kits for the MiSeq, which a commenter asked for more details. I've been remiss in not expanding on this sooner.
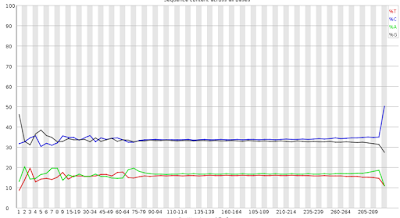
Here is part of a FASTQC plot for the first read of a 2x300 run where things went well. I've often been cavalier about running quality metrics on my data, letting my analyses be the only word, and what I'll illustrate has convinced me that this was not a good strategy. Getting back to the plot, what I am showing here is the portion that shows the fraction of the data that is A,C,G and T calls versus position. There is some noise at the front end, but then the lines stay relatively flat until the end. The separation between the G and C versus the A and T lines is indicative that we are sequencing wonderfully G+C rich Streptomycete genomes. As expected, the G and C lines are nearly coincident as are the A and T lines.
Here is R2 from that same sample. Note here that there is some separation of the G and C curves, but not the A and T. That's not good, and is reflected in a drop in the overall quality scores (not shown). It does make one wonder if this information could be incorporated further -- i.e. if instead of a simple basecall and quality score one could have a set of probabilities at each position for each possible base -- but that's a whole different can of worms. Note also that given the composition bias in my input data, this result strongly suggests Gs are being miscalled systematically as Cs (or vice versa; the color choices in FASTQC for those two are smack in the middle of my idiosyncratic color perception issues)
Okay, that's some not bad data, though I suspect if I skimmed through our wealth of 2x300 datasets I could probably find an even cleaner one. Now lets look at a dataset with major issues. Here's an R1 from summer 2015, showing more serious nucleotide composition artifacts, somewhat akin to the R2 above but even worse in the early going.
As you might expect, the R2 is truly horrible -- and we saw this with really terrible assembly results even with my favorite error-correcting assembler, SPADES
After this, and reports that one of the local university core facilities was simply refusing to run 2x300, we stopped using this mode. For a variety of reasons we haven't had cause to revisit that decision, so perhaps I'll wait for an all clear from the community. I did learn that ideally I should check these plots every time, and barring that should at least check them when things go south. Ideally my vendor would have been checking these, but (and no, I won't name&shame them; we generally have a good relationship) they didn't in this case.
I'm hardly the first person to point out issues with the 2x300 quality in general, and I believe that many core labs and MiSeq owners were aware of this issue, though there was only a little bit of Twitter talk about it. I suspect that these sorts of issues are also why we haven't seen any further improvements in read length, despite Illumina saying at user group meetings for several years now that they have demonstrated 2x400 or longer working. It also depends on your application; for sequence assembly of individual genomes aggressive error correction may be possible whereas for long amplicon sequencing with fusion primers you may be stuck with poor sequence quality where you most desire reliable sequence.



2 comments:
I am under the impression a lot of these problems are related to short insert size, so the whole reads + adapters are sequenced and then the machine starts to spout gibberish. On the lab I work, the suspicion / complain is library prep kits have changed, and average insert size is now 250bp whereas before was 400bp, using the same protocol.
To the previous commenter-
In more recent versions, the Illumina software trims the read if it detects the adapter, so those read throughs probably shouldn't show up in those QC charts. A couple of years ago this wasn't the case.
Post a Comment