Bradnam was, of course, kidding. His short item showered derision on a recent Microsoft announcment about importing sequences into Excel. Various twitterings in response to the article were along the same lines as Bradnam's punchline
The point at which you want to import FASTA files into Excel is the point at which you should probably think about quitting bioinformatics.
Thankfully, it didn't get quite as idiotic as the periodic bioinformatician who wears ignorance of Excel as a badge of honor.
My reaction? They're doing it wrong!
Microsoft Excel has a jaded reputation for bioinformatics, partly driven by snobbery against Microsoft and partly driven by real issues. Most notoriously, Excel's attempts to be helpful when importing comma or tab delimited files can be quite troublesome. Best known is the the issue of gene names which resemble dates (OCT1, MARCH1, etc) being mangled unless you explicitly import them as text. A issue that frequently crops up in my space is the practice of encoding wells on plates; if your use numerical barcodes separated from the well address with a period, then Excel will try to be helpful and convert 1234.E09 to 1,234,000,000,000. Also well noted is the fact that complex formulas in Excel are difficult-to-impossible to debug or validate.
Let's be clear here: I spend much of my time in Excel and typically have half a dozen or more worksheets open when my laptop finally crashes (Microsoft clearly has some memory leak issues!), but I also have a complex relationship with it. There are many "features" I would love to deep six, especially since with each new version they seem to drive the useful ones deeper and deeper into
the interface. Microsoft has an appalling affinity for glam and glitz, and horrific choices when it comes to defaults. I succumb to the temptation of plotting data in Excel about once a year ("it will be quicker"); between the awful defaults and the inability to do much exploring later, I quickly remember why I always insist on getting Spotfire.
But to dismiss Excel as unworthy of any use in bioinformatics is to miss the fact that buried under the residue of years of creeping featurism is a tool useful in specific contexts and with some key advantages. The first advantage is that it is ubiquitous, and not only is on all my colleagues' desktops but they all have at least a passing familiarity with it. Second, not only are few of my colleagues scared of Excel, but many actually know a few tricks for filtering or summarizing data, or at least for highlighting what is interesting to them. Plus they are comfortable with the multi-tabbed interface. Occasionally writing complex formulas in Excel is a gateway drug to serious programming; one of our RAs is going that route towards understanding SQL. These are opportunities!
The Microsoft post that Bradnam harshly spotlighted does classify as a fireable offense. In the few we see As, Ts, Cs and Gs in an Excel spreadsheet, Each cell is colored by what nucleotide it is, but there is no explanatory text beyond "Sequence data", so it isn't remotely clear what is going on here. Is this a single sequence? A multiple alignment? If a multiple alignment, are the individual sequences organized as rows or as columns? Without labels, this display is absolutely useless, but I'm not convinced the author of this image had any clue how to make things useful.
I've used Excel to deliver sequence analysis reports and will likely do so again in the future; I can say that without an ounce of sheepishness because I design my reports to leverage Excel and not be stymied by it or succumb to glitz.
As far as the data mangling upon import issue, this is easily fixed by not importing data via a delimited format, but rather using Excel's rich XML format. There are libraries out there for writing Excel, or it is easy to roll your own. By using the XML route, one can both prevent data mangling and format the data correctly (which I am very particular about) -- appropriate numbers of digits after the decimal (often zero, since so much bioinformatics is in integer space), coloring that actually means something, sequences in a monospace typeface, etc. The XML format also enables setting up multiple tabs. Even if you disdain importing sequences into Excel, understanding the XML system is an important skill for delivering oligo ordering forms -- IDT's (for example) allows you to designate each tab as a separate plate.
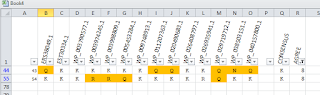
The first decision to make is which way sequences should run -- rows vs. columns. I've employed both, depending on how I might filter the data -- Excel has strong ability to filter rows but not columns. In the mock-up I threw together manually, I've used a single color to highlight discrepancies; if I had done this programmatically I might have gotten a little more extravagant -- though the danger of visual fruit salad is always present. But, when it makes sense, don't hesitate to use color -- as well as bold and italic -- to layer on information.
So, if I'm presenting a multiple alignment, I might display it vertically and then use additional columns for summary statistics. This enables my user to filter using the summary statistics, which might be a %conservation number -- in this mock-up I've filtered for the alignment "columns" (now rotated 90 degrees) which show the least conservation.
Alternatively, I might run the sequences horizontally if I think users might wish to filter the alignment given certain residues at certain positions. So, while I am showing all the rows in the alignment below, I could easily use Excel's filtering capability to show only the sequences with a specific deviation from the consensus -- or lack thereof. Alas, one can't in Excel filter the columns as well as rows, which would really make Excel a useful tool in this context.
I'd be the last person to claim that Excel should always be your method of delivering results -- only that you shouldn't rule it out. Its ubiquity and latent abilities combine to make it sometimes the right tool. Just because Microsoft is willing to be boldly and publicly stupid about a tool doesn't disqualify that tool.

1 comment:
I have found that delivering reports in Excel encourages wet-lab colleagues to explore the data themselves in ways they will not do with any other format. I do wish there were a global setting that told Excel, "always ASK before assuming gene names are dates." My favorite cases of gene names that get munged are MARC1, MARC1, MARCH1, and MARCH2: if Excel thinks DEC1 is a date, I can figure out what the original data file said but it turns both MARC1 and MARCH1 into the first of March in the current year.
Form Tools are a GREATLY underappreciated feature of Excel that enable really nice interactive reports. Below is an example I cooked up some years ago using data from the US Census website because most of the Excel tools I build have proprietary data from my workplace and I can't post those! Note that although I made this in an older version of Excel (hence the .xls file), it works fine in newer versions. Unfortunately doesn't work in Open Office or Libre Office:
http://mdhealy.home.sprynet.com/WorldPopulationInteractiveEstimates.xls
Post a Comment